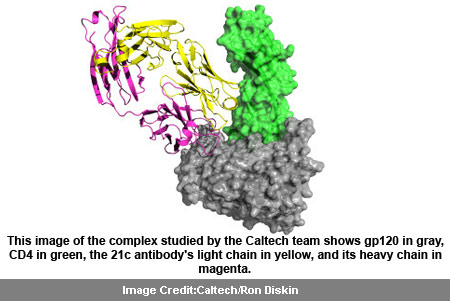
This 3D understanding of the construction of gp 120 seems to be more than just a basic scientific advancement. The image of the complex, studied by the team at Caltech identifies gp 120 in gray, CD4 receptor in green, the 21c antibody’s light chain in yellow, and its heavy chain in magenta.
Caltech postdoctoral scholar Ron Diskin says that, “There’s a tremendous continuing effort to develop a vaccine for HIV, and most of those efforts use gp120. Having more structural information will facilitate better vaccine design.â€
The team focused on gp 120 from what is known as clade C HIV-1. In order to understand the meaning of this, it is essential to know the family history of HIV. Most people who are affected with HIV are infected with the HIV-1 family of viruses. HIV-1 is segregated into different groups, and most of the AIDS-related symptoms are caused due to group M. These groups are further classified into something known as ‘clades’.
Clade B from group M HIV-1 family is the one that mostly attacks in the United States and Western Europe. This clade is supposedly the best analyzed clade till date. The clade studied by the Caltech team is clade C and has apparently terrorized Africa and Asia. It may be the one that is causing the largest number of infections globally. Previous studies that looked into the structure of clade B gp 120, assume that clade C might share a more or less similar structure.
In order to uncover the structure of clade C gp 120 and to understand if the similarities do really exist, the team had to crystallize the protein. However the crystallization process did not go smoothly as the protein was not stiff enough. Hence, the scientists created a complex of molecules comprising of gp 120 monomer, a CD4 receptor and an anti-HIV antibody known as 21c.
This complex helped with the crystallization process thereby allowing the scientists to look at not only gp 120 but also to visualize the entire binding site and to understand the interaction of the various elements within the complex. The team did find resemblance between clade C and clade B.
The scientists during this study realized that the Antibody 21c was not only reactive to the gp 120 protein sticking out of the HIV envelope, but was also reacting to the CD4 receptor present in the body’s own T cells. It is apparently for the first time that this sort of polyreactivity, i.e. a response to multi-antigens, had been noticed in the 3D structure of the HIV- targeted antibody.
Pamela Bjorkman, the Max Delbruck Professor of Biology at Caltech, a Howard Hughes Medical Institute investigator, and the Caltech team’s leader, said that, “The most interesting aspect of our structure is the unexpected contact between the antibody and CD4. The binding to CD4 suggests that this class of anti-HIV antibodies has autoreactive properties, which raises many interesting questions about how anti-HIV immune responses affect an HIV-infected individual.â€
This gives rise to the question that does autoreactivity symbolize that 21c is too dangerous to work with, as clinicians might be facing potential autoimmune response with a vaccine that elicits 21c like antibodies?
Diskin replies to the question by stating that, “Not necessarily. Other data out there show that some of the best neutralizing antibodies are also autoreactive.â€
Diskin goes on to explain, that what the autoreactivity actually means is that there could be additional hurdles to overcome in eliciting such antibody responses. The body tends to get rid of autoreactive antibodies with the sole purpose of keeping autoimmune diseases away. The study shows that to create an effective vaccine to produce 21c like antibodies, scientists would need to comprehend ways of overcoming this elimination process.
The follow-on for the Caltech team is to work on the relatively low resolution structure. Nevertheless, the team will also be trying to clear the structure of gp 120 trimer, which is a more complex, three-pronged version of the protein.
The findings of this study were published in the advance online edition of the journal Nature Structural & Molecular Biology.
