This latest study may indicate one of the first large-scale reconstructions of a mammalian circuit and may propose a practical approach for unraveling the circuits that apparently support other significant biological systems.
Senior author Nir Hacohen, of the Massachusetts General Hospital (MGH) Center for Immunology and Inflammatory Diseases, an assistant professor at Harvard Medical School and a senior associate member at the Broad Institute, commented, “Our findings address a fundamental question in human biology: how do immune cells recognize various pathogens and use that information to mount distinct responses. We now have a detailed view of the circuitry that controls this critical process, providing a deeper understanding of immune biology that could inspire novel ways to treat disease and design better vaccines.â€
Cells may receive and process information much like computers. Information pours in, is studied and processed through an intricate set of circuits, and a suitable response may be conveyed. But instead of minute transistors, the internal circuitry of mammalian cells is apparently made up of immense networks of genes and their equivalent proteins. A frontier of modern genomic research is to supposedly recognize these molecular parts and their interconnections, which may replicate the usual and occasionally faulty ‘wiring’ that may cause human biology and disease. Until lately, study in this region claimed to focus on yeast and bacteria as it seemed virtually unfeasible to carry it out in mammals.
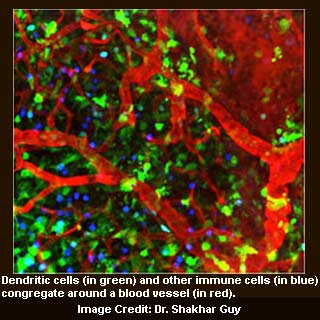
Since there was a lot of interest in the specialized immune cells called as dendritic cells, the experts apparently wanted to expose the full scope of their internal circuitry. Dendritic cells are claimed to be among the first to detect pathogens and may distinguish one kind of pathogen from another, supposedly letting them to arrange a pathogen-specific immune response. These cells may achieve such tasks through two critical functions. The first function is that they may offer tiny pieces of an infecting pathogen to other immune cells so those cells can learn to identify it. The second is they may initiate other cells to respond in ways that could eradicate the culprit pathogen.
Lead author Ido Amit, a postdoctoral fellow at the Broad Institute and Massachusetts General Hospital, along with his colleagues assembled a detailed depiction of the circuit’s output. This was done by gauging the activities of thousands of genes after mixing parts of diverse pathogens, including RNA from viruses and pieces of the bacteria Escherichia coli, with primary mouse dendritic cells.
Subsequently, they battered these data to make out genes whose activities may alter with varying situations. These genes could form the circuit’s potential regulatory framework and may be accountable for controlling the course of information.
To find out how these potential regulators work and how they are wired together, the researchers apparently methodically eradicated each of them, and recorded the changes in the circuit’s output. This was apparently made possible by the use of RNA interference (RNAi), which may decrease or ‘knock down’ the activity of particular genes and may be applied to basically every gene in the mouse genome. A single-molecule technology apparently allowed the experts to generate sensitive readouts of gene activity.
Regev, Hacohen, Amit and their colleagues exposed a dendritic cell circuit with two major arms i.e. an inflammatory arm, which may be highly active during bacterial infections and may begin a system-wide immune response; and an anti-viral arm, which could persuade upon viral infections and apparently synchronizes a more focused response tailored to viruses. Together, these arms may encompass about 100 regulators about four times as many as were earlier known to be involved and could contain several proteins that were not assumed to direct immune responses.
The experts apparently recognized an astonishing number of connections between regulators and other circuit components, approximately more than 2,300 connections in total. In addition, some regulators appear to control a relatively broad swath of the circuit, including 25 genes or more, while others may sway just a handful of genes.
One captivating ‘coarse tuner’ is a protein known as Timeless. In fruit flies, it controls circadian rhythms, the internal clock that may keep biological processes working on a 24-hour cycle. In mammalian dendritic cells, it was discovered that Timeless may be a chief regulator of anti-viral responses, supposedly controlling over 200 genes required to combat viruses.
Another remarkable regulator is CBX4, a ‘fine tuner’ that may control the levels of a key protein caught up in viral infections. This protein, known as IFNB1 (for Interferon beta 1) needs accurate control. If a virus is present, it may be highly active, yet if bacteria are the offending agents, its activity should apparently be curtailed.
While the findings may basically be mechanistic in nature and do not yet have any direct bearing on human disease, there may be a few imperative medical allegations. For example, an absolute understanding of the regulatory network that may control immune responses may assist in laying the groundwork for more accurate involvement, including drugs and vaccines.
Additionally, 12 of the regulators apparently recognized in the Science study exist in areas of the genome that may have been connected to autoimmune and other associated diseases in humans, and may boost the understanding of the genetic susceptibilities to infections and other immune disorders.
This study was published in the journal Science.
